Tandem Shady Sayed & Carina Wenzel
Next-Generation Precision Oncology: Genome Editing in Translational Lung Cancer Research
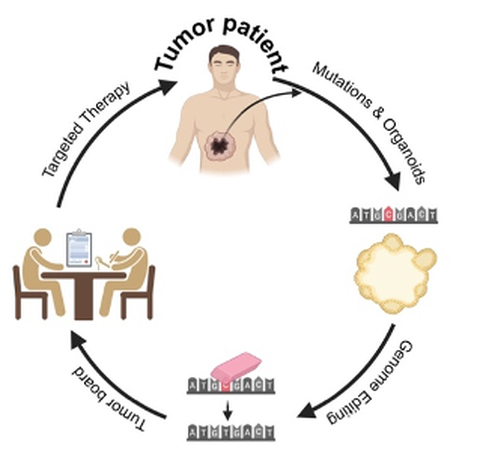
Lung cancer remains the leading cause of cancer-related mortality in the European Union, with approximately 249,000 deaths each year. More than half of patients are diagnosed with metastatic disease, necessitating systemic, drug-based therapies. Over the past decade, systemic treatment options, including the use of targeted therapies for tumors with specific mutations have expanded rapidly. However, given the low incidence of such mutations, only the minority of lung cancer patients are eligible for these treatments. Furthermore, the use of these drugs almost always leads to the development of resistances, which diminishes their long-term efficacy. Meanwhile, the precise correction of mutant TP53 in its endogenous context offers a powerful strategy to restore its tumor-suppressive function. Up to 70% of lung tumors carry a TP53 mutation. Therefore, our research focuses on modeling patient-specific mutations, especially TP53 mutations, by applying CRISPR-based mutation repair and functional analysis in patient-derived lung tumor organoids to profile oncogenic mutations and identify mutation-driven therapeutic vulnerabilities.
Our previous work have demonstrated that CRISPR-based genome editing can not only uncover previously unrecognized cancer-driving mutations, but also precisely correct oncogenic variants within their native genomic environment, resulting in loss of tumor cell viability. Moreover, this approach enables the functional stratification of mutations in key cancer driver genes, e.g. TP53, KRAS, and SMAD4, based on their impact on cellular fitness and tumor dependency.
To enhance clinical translation, we are expanding these studies into patient-derived model systems, particularly lung tumor organoids, which preserve the genetic, phenotypic, and therapeutic heterogeneity of primary tumors. These models allow us to evaluate genome-editing–based mutation repair in a clinically relevant context and to assess its impact on tumor growth, survival, and therapeutic response.
Current efforts further focus on improving tumor- and tissue-specific delivery of genome editing components using advanced nanocarrier platforms, thereby increasing feasibility for in vivo applications. By integrating genome editing, nanocarrier-based delivery, and lung tumor patient-derived organoid models, our mutation-repair strategy establishes a translational framework for precision oncology. Ultimately, this approach is designed to serve as a scalable blueprint for the development of mutation-targeted therapeutic strategies across multiple cancer types.
Research aims
- Functional characterisation of oncogenic mutations in lung cancer following mutations repair, to guide more effective personalised therapies.
- Screen nanocarrier delivery vehicles e.g. lipid nanoparticles, across a panel of cancer organoids to profile cell and tumor-type specificity.
- In vivo molecular surgery utilising selected CRISPR systems to repair oncogenic mutations as a therapeutic modality in mouse models of cancer.
Members
Dr. Shady Sayed, Junior Group Leader MSNZ
Dr. Carina Wenzel, Junior Group Leader MSNZ
Solveig Freda Klier, Master Student
Jean Michel Beutler, Bachelor Student
Selected publications
Pascal Wang, Rituparno Sen, Frank Buchholz, and Shady Sayed. A base editing platform for the correction of cancer driver mutations unmasks conserved p53 transcription programs. Genome Biol, Jul 2025. https://doi.org/10.1186/s13059-025-03667-7. The publication was awarded “Paper of the Quarter” by the German Society for Gene Therapy, https://www.dg-gt.de/paper-of-the-quarter
Lukas Schmitt, …, Shady Sayed, Frank Buchholz, and Duran Sürün. Quantification of evolved DNA-editing enzymes at scale with DEQSeq. Genome Biol, Nov 2023. 10.1186/s13059-023-03097-3
Shady Sayed, …, and Frank Buchholz. Efficient Correction of oncogenic KRAS and TP53 mutations through CRISPR Base Editing. Cancer Res, Sept 2022. 10.1158/0008-5472.CAN-21-2519
Shady Sayed, …, and Frank Buchholz. Using CRISPR-Cas9 to Dissect Cancer Mutations in Cell Lines. Cancer Cell Biology. Methods Mol Bio, vol 2508. Jun 2022. 10.1007/978-1-0716-2376-3_18
Shady Sayed, Maciej Paszkowski-Rogacz, …, and Frank Buchholz. CRISPR/Cas9 as a tool to dissect cancer mutations, Methods, May 2019. 10.1016/j.ymeth.2019.05.007
Frost N, Joosten M, Franzen J, Wiesweg M, Rasokat A, Kulhavy J, Kollmeier J, Reinmuth N, Grohé C, Roeper J, Rittmeyer A, Heinzen S, Wermke M, Wesseler C, Christopoulos P, Kauffmann-Guerrero D, Althoff A, Bleckmann A, Collienne M, Berezucki E, Overbeck T, Kropf-Sanchen C, Griesinger F, Sebastian M, Schuler M, Braun S, Wenzel C, Furth C, Wolf J, Bischoff P, Reck M; National Network Genomic Medicine Lung Cancer Germany. PET/CT-guided management of immune checkpoint blockade and multi-modal profiling following treatment in long-term responders with metastatic lung cancer in the National Network Genomic Medicine Lung Cancer Germany (nNGM). Ann Oncol. 2025 Dec 30:S0923-7534(25)06329-X. doi: 10.1016/j.annonc.2025.12.011. Epub ahead of print. PMID: 41478526.
Kropf-Sanchen C, Rasokat A, Christopoulos P, Wenzel C, Wehler T, Rost M, Kulhavy J, Reinmuth N, Schulz C, Scheffler M, Wolf J, Büttner R, Merkelbach-Bruse S, Thomas M, Stenzinger A, Schütz M, Bräuninger A, Demes M, Hummel HD, Pfarr N, Gaisa NT, Rawluk J, Berezucki E, Lutz KT, Galda S, Jacobi H, Collienne M, Janning M, Brummer T, Loges S; nNGM Lung Cancer. Treatment outcome of NSCLC patients with BRAFnon-V600E mutations: a retrospective, multicentre analysis within the national Network Genomic Medicine (nNGM) Lung Cancer in Germany. ESMO Open. 2025 Aug;10(8):105124. doi: 10.1016/j.esmoop.2025.105124. Epub 2025 Jul 14. PMID: 40664146; PMCID: PMC12281862.
Saalfeld FC, Möller J, Christopoulos P, Wenzel C, Rasokat A, Wang XA, Vathiotis I, König D, Illini O, Grohé C, Wiesweg M, Wesseler C, Schubart C, Pelusi N, Rohde G, Overbeck TR, Kirfel J, Alt J, Kauffmann-Guerrero D, Griesinger F, Kulhavy J, Allgäuer M, Klimova A, Schütz M, Aust DE, Hochmair MJ, Rothschild SI, Syrigos KN, Veluswamy R, Michels S, Stenzinger A, Jöhrens K, Wermke M. Small cell transformation in EGFR-mutated non-small cell lung cancer: DLL3 expression and efficacy of immune checkpoint inhibitors or tyrosine kinase inhibitors combined with chemotherapy. Eur J Cancer. 2024 Dec;213:115065. doi: 10.1016/j.ejca.2024.115065 Epub 2024 Oct 12. PMID: 39423775.
Schmäche T, Fohgrub J, Klimova A, Laaber K, Drukewitz S, Merboth F, Hennig A, Seidlitz T, Herbst F, Baenke F, Ada AM, Groß T, Wenzel C, Ball CR, Praetorius C, Schmidt T, Ringelband-Schilling B, Koschny R, Stenzinger A, Roeder I, Jaeger D, Zeissig S, Welsch T, Aust D, Glimm H, Folprecht G, Weitz J, Haag GM, Stange DE. Stratifying esophago-gastric cancer treatment using a patient-derived organoid-based threshold. Mol Cancer. 2024 Jan 10;23(1):10. doi: 10.1186/s12943-023-01919-3. PMID: 38200602; PMCID: PMC10777586.
Wenzel C, Herold S, Wermke M, Aust DE, Baretton GB. Routine Molecular Pathology Diagnostics in Precision Oncology. Dtsch Arztebl Int. 2021 Apr 16;118(Forthcoming):255–61. doi: 10.3238/arztebl.m2021.0025. Epub ahead of print. PMID: 33536117; PMCID: PMC8287073.