Jan 08, 2024
New Publication out !
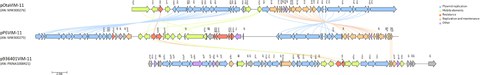
The image shows the linear maps of blaVIM-11 harboring plasmids. Plasmids were recovered from P. aeruginosa in Argentina.
Pseudomonas aeruginosa is a major cause of hospital-acquired infections, with high morbidity-mortality rates linked to resistant phenotypes. Class B metallo-β-lactamases (MBL) are prevalent in this species, with VIM being the most frequent. VIM-11, a widespread MBL, has better hydrolytic efficiency for imipenem and slightly better catalytic efficiency for ceftazidime, cefepime, and cefpirome than VIM-2. blaVIM-11, has been described as a gene cassette in the variable region of class 1 integrons in P. aeruginosa and Enterobacterales but no records of such genetic structure were publicly available. Here we report the first full descriptions of blaVIM-11 harbouring plasmids recovered from human-infecting P. aeruginosa.
Elena AX, Cejas D, Gutkind G, Radice, M (2023) Genomic characterization of blaVIM-11 harboring plasmids recovered from Pseudomonas aeruginosa. Journal of Global Antimicrobial Resistance. doi.org/10.1016/j.jgar.2023.12.014
~~~
more publications here ...
