The Antos Group
My group is interested in tissue regeneration, including cardiac physiology. To understand the biology of regeneration, I am interested in answering three fundamental questions:
1. How are cells at the site of injury induced to regenerate lost tissues.
2. How are cells involved in regeneration controlled to produce the correct pattern.
3. What stops regeneration once the appropriate size is reached.
My group uses the zebrafish to answer these questions, because the zebrafish will regenerate heart, appendages, etc. Therefore, we can describe and dissect the cell and molecular mechanisms involved in the fish’s regeneration abilities.
Healthy heart physiology requires coordinated Ca2+ handling across the syncytial cardiac muscle. While single-cell cardiomyocyte experiments in culture indicate how individual cardiomyocytes alter their Ca2+ subcellular distribution and handling properties, they are limited in addressing the syncytial nature of the cardiomyocytes in the heart in vivo. Thus, experimental dissection at the organ level is needed to understand how known (catecholamine and natriuretic signalling) and to-be-discovered mechanisms regulate cardiac physiology in vivo. We are therefore using real-time in vivo zebrafish heart imaging to address fundamental questions about cardiac contractility, injury and regeneration.
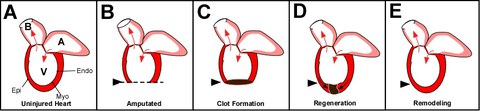
Figure 1: Zebrafish heart regeneration. A) The adult zebrafish heart consists of one ventricle (V), one atrium (A) and the balbus artieriosis (B). Arrows indicate the direction of blood flow. Similar to mammalian heart walls, the fish has an epicardium (Epi), myocardium (Myo) and endocardium (Endo). The difference between mammalian adult heart and the zebrafish adult heart is that the fish regenerates injury. Amputation of the ventricular wall (B) will result in the formation of a blood clot (C). This clot and subsequent scar tissue will be resolved as new myocardium, epicardium and endocardium forms and replaces the scar tissue (D) until it has regenerated (E).
Determine how the calcineurin is integrated into the biology regulating proportional growth.
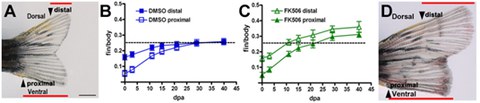
This research focus relates to two questions: 1) what initiates regenerative outgrowth and 2) what stops regeneration at the proportionally correct size. Inhibition of calcineurin activity results in fins that regenerate beyond their original dimensions due to an extended higher regenerative growth rate. The rate of regeneration is coupled to proximodistal position in the appendage. My lab showed that inhibition of calcineurin results in fins that regenerate beyond their original dimensions due to a maintained proximal growth rate (Fig. 2). Furthermore, inhibition of calcineurin in uninjured fins induces outgrowth. Thus, determining the regulation of Ca2+ and calcineurin, the mechanism that calcineurin regulates, and the cells in which this occurs will lead to subsequent hypothesis-based exploration of the cellular and molecular relationships between positional information and proportional growth as well as the promotion of coordinated outgrowth.
Figure 2: Calcineurin regulates positionally determined growth rate. (A) Amputations of one lobe proximally cut (proximal arrow) and one distally cut (distal arrow). (B) Graph shows the relationship in growth rates between proximally and distally cut regenerating lobes in relation to the body length and show the reduction of their rates as the fin regenerates to the original size of each lobe. (C) Inhibition of calcineurin maintains both regulating lobes at a proximal growth rate so that they continue to grow in parallel beyond the original dimensions of the fin. (D) Each lobe regenerates at the same proximal rate after calcineurin inhibition, so the lobes do not converge [Adapted from Kujawski et al., 2014].
By quantitative modeling of in vivo Ca2+ distribution patterns in specific tissues, regions of tissues and subcellular domains during regeneration, we will look for how Ca2+regulates calcineurin to prevent allometric growth using standard fluorescent confocal microscopy. I hypothesize that the intracellular Ca2+-signal is localized to nanodomains in which calcineurin acts to suppress growth, and we are creating transgenic calcium reporter lines to identify these domains. Once identified, we will be able to link these domains with the channels and the signals that regulate them. We can then define the origin of these signals; and thereby, continue to explore the Ca2+-calcineurin-linked mechanisms regulating proportional growth. Since Ca2+ is a second messenger to several enzymes, a broader knowledge of subcellular Ca2+ dynamics during tissue regeneration will lead to other avenues of research exploring its potential in vivo roles in progenitor cell formation, migration, differentiation in addition to growth control. We are also pursuing the mechanisms that promote growth by looking for the reciprocal kinase(s) and by defining the downstream calcineurin-regulated mechanisms. From existing zebrafish mutants, the scientific literature and deep sequencing results, we already have candidate targets and mechanisms to test: a potassium channel, a connexin, transcription factors, growth factors and a chromatin remodeling enzyme. Furthermore, the discovery that fin regenerative outgrowth can be extended by drugs means that we can perform small molecule and chemical screens to identify drug candidates and other components of the mechanisms regulating proportional growth control and set up a background mutagenesis screen for fin growth mutants.
Determine how Smp regulates β-catenin-dependent Wnt signaling and whether it has other functions.
My group identified simplet/fam53b (smp) expression in the progenitor cells of regenerating hearts, livers and fins from transcriptome bioinformatic analyses. This gene is conserved among vertebrate species, but its function was unknown. We showed that smp is required to regulate cell proliferation and tissue patterning during fin regeneration. Thus, studying smp function will allow me to understand the mechanisms in regenerative growth and patterning.
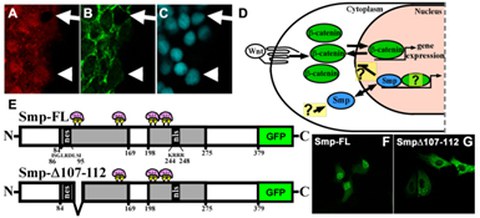
Figure 3: Smp is required for β-catenin-dependent Wnt signaling. (A) Immunohisto-chemistry stains for endogenous Smp, (B) for β-catenin, and (C) for DAPI. Arrows show absence of nuclear Smp and β-catenin. Arrowheads show nuclear colocalization of Smp and β-catenin. (D) Hypothesized mechanisms and questions on how Smp is regulated and how it regulates β-catenin and other partners. (E) Smp protein diagrams showing the two conserved homology regions (HR) (grey) and the nuclear localization signal in the second HR (nls) and a nuclear export signal (nes) in the first HR. (F) Smp is primarily localized in the nucleus with some cytoplasmic localization. (G) Overexpression of the deletion mutant in (E) results in complete cytoplasmic localization of the protein.
My group subsequently found that Smp regulates β-catenin nuclear localization, that this regulation requires the nuclear localization signal in the Smp protein, that Smp regulates β-catenin by retaining it in the nucleus, and that this protein is present in the stem/progenitor cells of mammalian intestinal crypts, another site of canonical Wnt signaling. Canonical Wnt signaling is important for stem cell maintenance, developmental patterning, and tissue regeneration, and its regulating by other signal transduction mechanisms is necessary for the controlled growth and patterning. Our discoveries that Smp regulates β-catenin nuclear localization and that Smp nuclear localization is independent of Wnt signaling as well as our evidence that Smp shuttles between the cytoplasm and nucleus all argue that Smp is a regulatory node between β-catenin and other signaling mechanisms (Fig. 3A-D). Furthermore, we recently identified sequences in Smp that regulate its nucleus-cytoplasm shuttling (Fig. 3E-G). Therefore, the questions we will address are what regulates the subcellular distribution of Smp, how exactly does Smp regulate β-catenin nuclear localization, and whether Smp interacts with other nuclear proteins to regulate gene transcription independent of β-catenin. To answer these questions, we will test candidate β-catenin regulators, identify other interaction partners by mass spectrometry, and assess how post-translational modifications regulate Smp subcellular distribution. By measuring the real-time shuttling Smp and β-catenin in developing zebrafish embryos and regenerating fins and measuring β-catenin transcriptional activity using transgenic zebrafish reporter lines, we can assess and model the dynamics of β-catenin regulation during tissue development and regeneration. By identifying how other signaling pathways regulate Smp, we will learn about the post-transcriptional regulation of β-catenin and other nuclear factors. One of which is 14-3-3. Thus, my discoveries have broader implications in research fields of embryonic development and cancer due to the importance of β-catenin-dependent signaling in these phenomena.